Simulates from a conditional bivariate copula, where each copula parameter takes a different value, depending on the calibration function and covariates.
condBiCopSim(family, calib.fnc, X, par2 = 0, return.par = TRUE, tau = TRUE)Arguments
- family
family A copula family:
1Gaussian,2Student t,3Clayton,4Gumbel,5Frank,13Survival Clayton,14Survival Gumbel,23Rotated (90 degrees) Clayton,24Rotated (90 degrees) Gumbel,33Rotated (270 degrees) Clayton and34Rotated (270 degrees) Gumbel.- calib.fnc
A calibration function.
- X
A vector (if
calib.fnctakes a single argument) or matrix (ifcalib.fnctakes multiple arguments) of covariates values.- par2
The second copula parameter (for the Student t), default
par2 = 0.- return.par
Should the parameter (and calibration function) be returned as well (default
return.par = TRUE)?- tau
Should the calibration function (and the model) be specified for the copula parameter or Kendall's tau (default
tau = TRUE)?
Value
If return.par = TRUE, then the function returns a list with:
data, a matrix with two columns containing the simulated data,par, a vector containing the values of the copula parameter,and
eta, a vector containing the values of the calibration function.
If return.par = FALSE, then the function simply returns data,
a matrix with two columns containing the simulated data.
See also
Examples
require(copula)
#> Loading required package: copula
set.seed(0)
## Simulation parameters (sample size, correlation between covariates,
## Gaussian copula family)
n <- 2e2
rho <- 0.5
fam <- 1
## A calibration surface depending on three variables
eta0 <- 1
calib.surf <- list(
calib.quad <- function(t, Ti = 0, Tf = 1, b = 8) {
Tm <- (Tf - Ti) / 2
a <- -(b / 3) * (Tf^2 - 3 * Tf * Tm + 3 * Tm^2)
return(a + b * (t - Tm)^2)
},
calib.sin <- function(t, Ti = 0, Tf = 1, b = 1, f = 1) {
a <- b * (1 - 2 * Tf * pi / (f * Tf * pi +
cos(2 * f * pi * (Tf - Ti))
- cos(2 * f * pi * Ti)))
return((a + b) / 2 + (b - a) * sin(2 * f * pi * (t - Ti)) / 2)
},
calib.exp <- function(t, Ti = 0, Tf = 1, b = 2, s = Tf / 8) {
Tm <- (Tf - Ti) / 2
a <- (b * s * sqrt(2 * pi) / Tf) * (pnorm(0, Tm, s) - pnorm(Tf, Tm, s))
return(a + b * exp(-(t - Tm)^2 / (2 * s^2)))
}
)
## Display the calibration surface
par(mfrow = c(1, 3), pty = "s", mar = c(1, 1, 4, 1))
u <- seq(0, 1, length.out = 100)
sel <- matrix(c(1, 1, 2, 2, 3, 3), ncol = 2)
jet.colors <- colorRamp(c(
"#00007F", "blue", "#007FFF", "cyan", "#7FFF7F",
"yellow", "#FF7F00", "red", "#7F0000"
))
jet <- function(x) rgb(jet.colors(exp(x / 3) / (1 + exp(x / 3))),
maxColorValue = 255
)
for (k in 1:3) {
tmp <- outer(u, u, function(x, y)
eta0 + calib.surf[[sel[k, 1]]](x) + calib.surf[[sel[k, 2]]](y))
persp(u, u, tmp,
border = NA, theta = 60, phi = 30, zlab = "",
col = matrix(jet(tmp), nrow = 100),
xlab = paste("X", sel[k, 1], sep = ""),
ylab = paste("X", sel[k, 2], sep = ""),
main = paste("eta0+f", sel[k, 1],
"(X", sel[k, 1], ") +f", sel[k, 2],
"(X", sel[k, 2], ")",
sep = ""
)
)
}
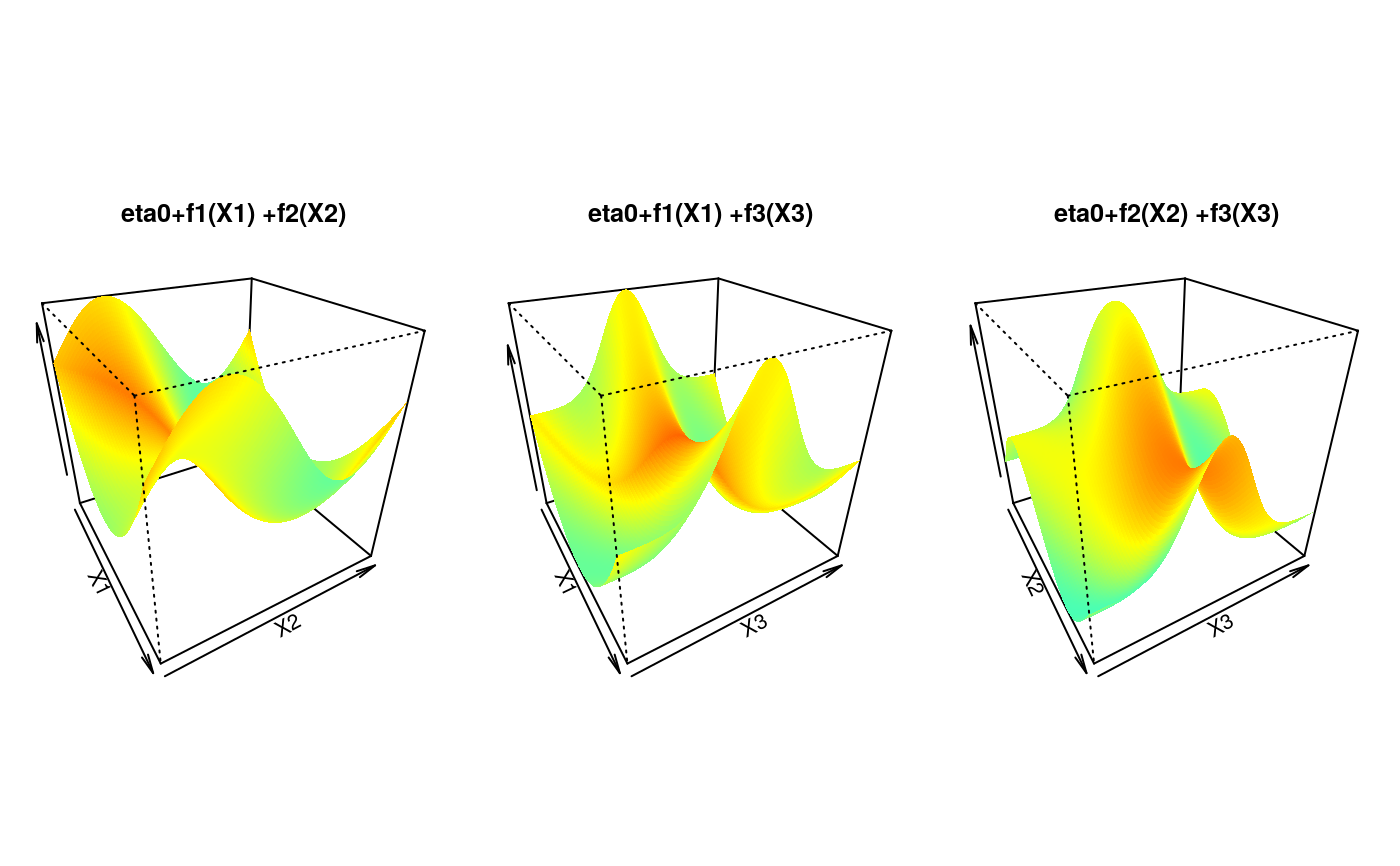 ## 3-dimensional matrix X of covariates
covariates.distr <- mvdc(normalCopula(rho, dim = 3),
c("unif"), list(list(min = 0, max = 1)),
marginsIdentical = TRUE
)
X <- rMvdc(n, covariates.distr)
## U in [0,1]x[0,1] with copula parameter depending on X
U <- condBiCopSim(fam, function(x1, x2, x3) {
eta0 + sum(mapply(function(f, x)
f(x), calib.surf, c(x1, x2, x3)))
}, X[, 1:3], par2 = 6, return.par = TRUE)
## Merge U and X
data <- data.frame(U$data, X)
names(data) <- c(paste("u", 1:2, sep = ""), paste("x", 1:3, sep = ""))
## Display the data
dev.off()
#> null device
#> 1
plot(data[, "u1"], data[, "u2"], xlab = "U1", ylab = "U2")
## 3-dimensional matrix X of covariates
covariates.distr <- mvdc(normalCopula(rho, dim = 3),
c("unif"), list(list(min = 0, max = 1)),
marginsIdentical = TRUE
)
X <- rMvdc(n, covariates.distr)
## U in [0,1]x[0,1] with copula parameter depending on X
U <- condBiCopSim(fam, function(x1, x2, x3) {
eta0 + sum(mapply(function(f, x)
f(x), calib.surf, c(x1, x2, x3)))
}, X[, 1:3], par2 = 6, return.par = TRUE)
## Merge U and X
data <- data.frame(U$data, X)
names(data) <- c(paste("u", 1:2, sep = ""), paste("x", 1:3, sep = ""))
## Display the data
dev.off()
#> null device
#> 1
plot(data[, "u1"], data[, "u2"], xlab = "U1", ylab = "U2")